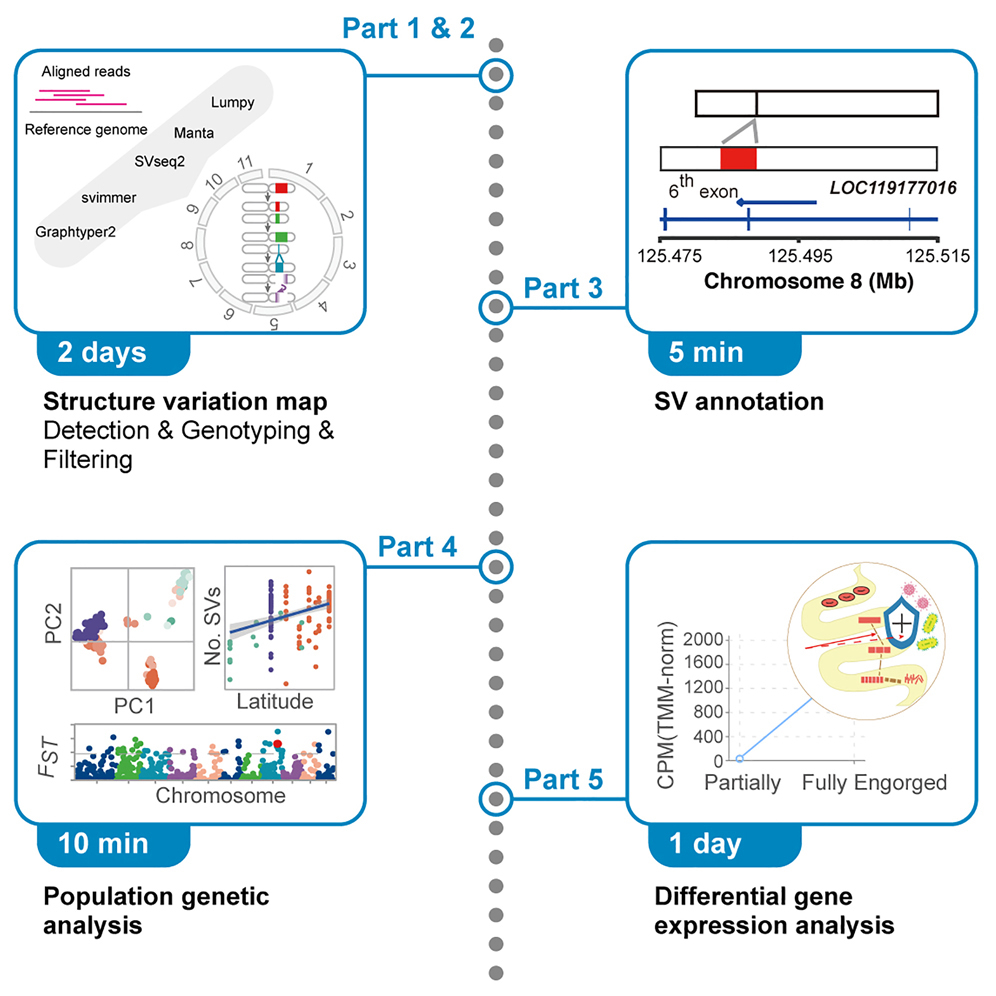
GitHub - tgac-vumc/ACE: Absolute Copy Number Estimation using low-coverage whole genome sequencing data
By A Mystery Man Writer
Description
Absolute Copy Number Estimation using low-coverage whole genome sequencing data - tgac-vumc/ACE

Accucopy: accurate and fast inference of allele-specific copy number alterations from low-coverage low-purity tumor sequencing data, BMC Bioinformatics

PDF) Genome‑wide copy number analysis of circulating tumor cells in breast cancer patients with liver metastasis

GATK-gCNV: A Rare Copy Number Variant Discovery Algorithm and Its Application to Exome Sequencing in the UK Biobank
GitHub - AdelmanLab/GetGeneAnnotation_GGA

Low-Pass Genome Sequencing: Validation and Diagnostic Utility from 409 Clinical Cases of Low-Pass Genome Sequencing for the Detection of Copy Number Variants to Replace Constitutional Microarray - ScienceDirect
Bamgineer: Introduction of simulated allele-specific copy number variants into exome and targeted sequence data sets

RNAseqCNV: analysis of large-scale copy number variations from RNA-seq data
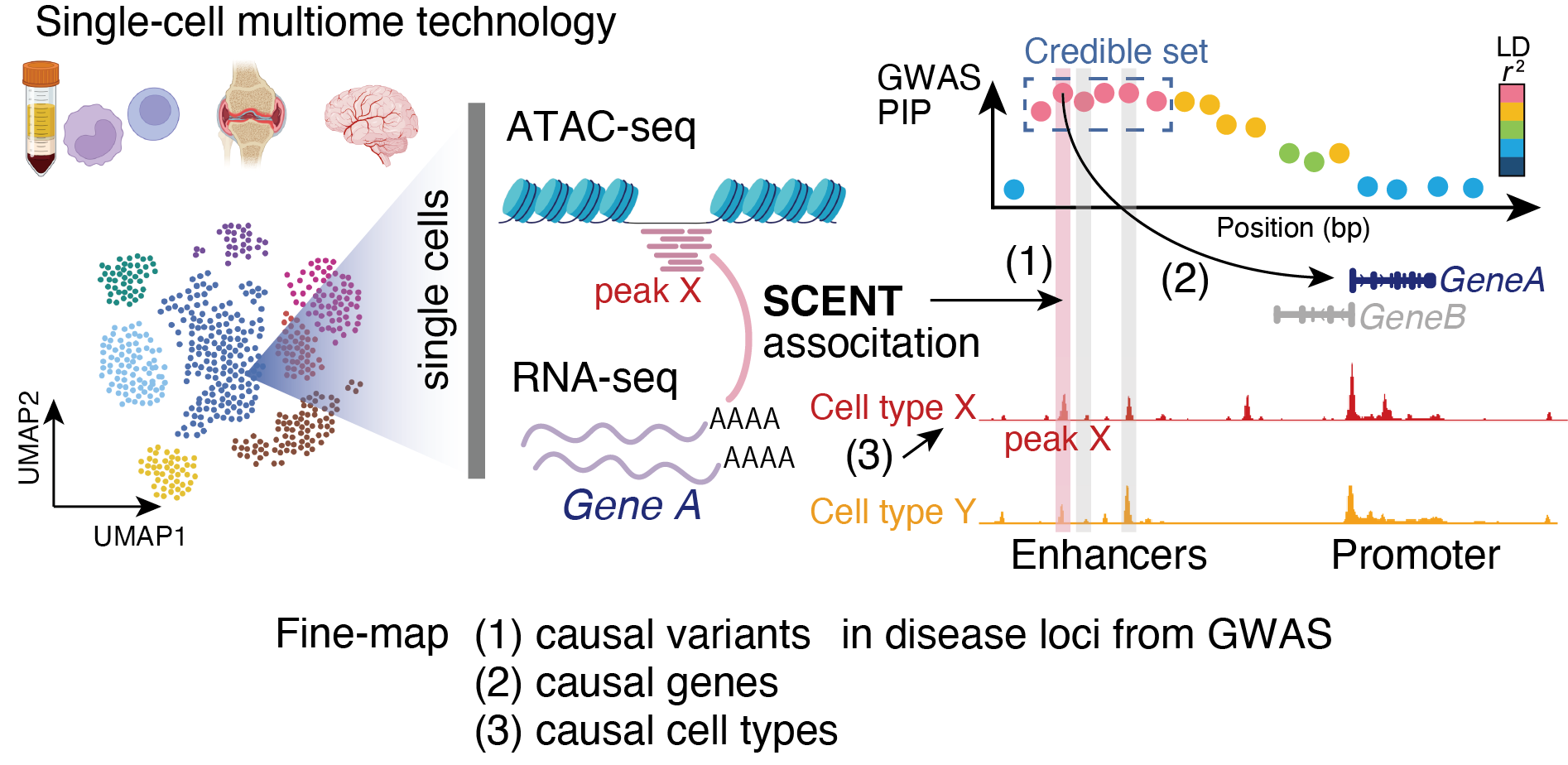
GitHub - immunogenomics/SCENT: Single-Cell ENhancer Target gene mapping using multimodal data with ATAC + RNA

PDF) PCR-Free Shallow Whole Genome Sequencing for Chromosomal Copy Number Detection from Plasma of Cancer Patients Is an Efficient Alternative to the Conventional PCR-Based Approach

Copy number analysis by low coverage whole genome sequencing using ultra low-input DNA from formalin-fixed paraffin embedded tumor tissue, Genome Medicine

Copy Number Variant Detection Using Next-Generation Sequencing - ScienceDirect

Shallow whole-genome sequencing of plasma cell-free DNA accurately differentiates small from non-small cell lung carcinoma, Genome Medicine
from
per adult (price varies by group size)