Technical spotlight: Detecting small- and medium-length copy number variants by whole-genome sequencing
By A Mystery Man Writer
Description
Historically, detecting different sizes of genetic variants has required using multiple different tests. By combining Illumina WGS with secondary analysis algorithms built into the DRAGEN Bio-IT Platform, researchers can achieve high-sensitivity detection of all these different variant types using a mixture of methods described here.

Identification of Copy Number Alterations from Next-Generation Sequencing Data
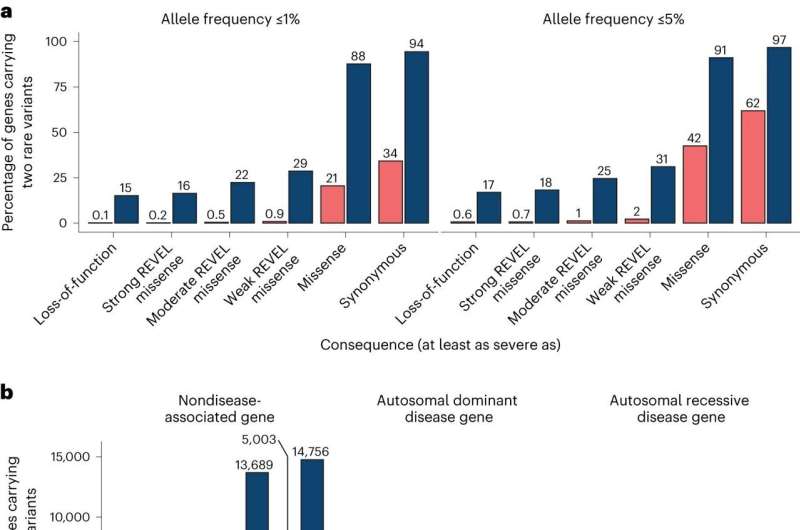
Researchers develop approach to study rare gene variant pairs that contribute to disease

Genomics Research Illumina research & innovation
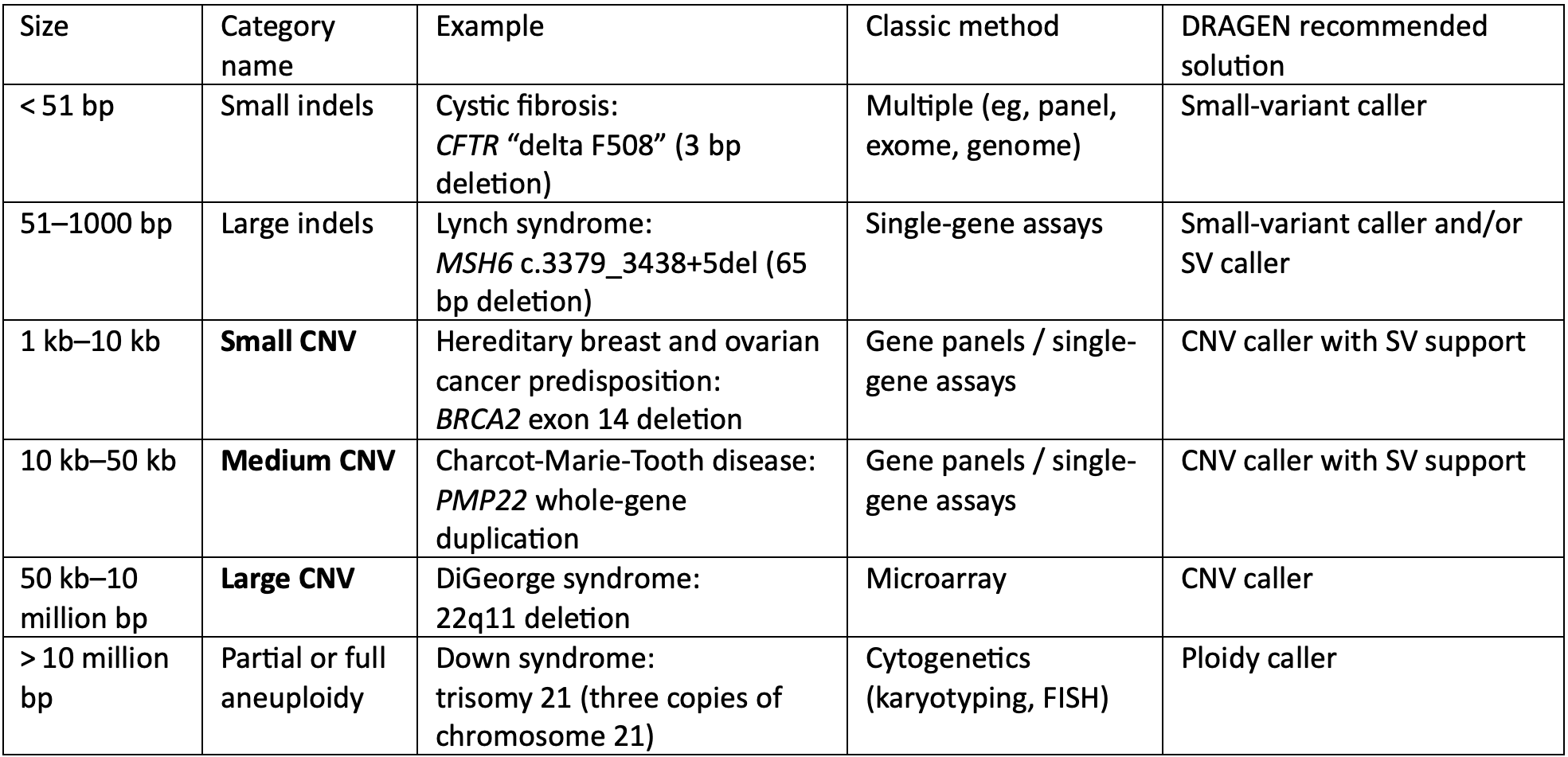
Technical spotlight: Detecting small- and medium-length copy number variants by whole-genome sequencing
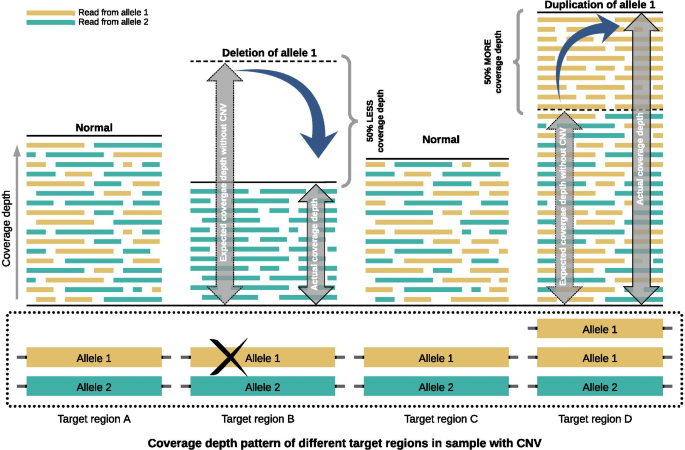
Detecting copy number variation in next generation sequencing data from diagnostic gene panels, BMC Medical Genomics
Single-cell copy number variant detection reveals the dynamics and diversity of adaptation

SV detection strategies using short reads. (A) Read-depth-based method
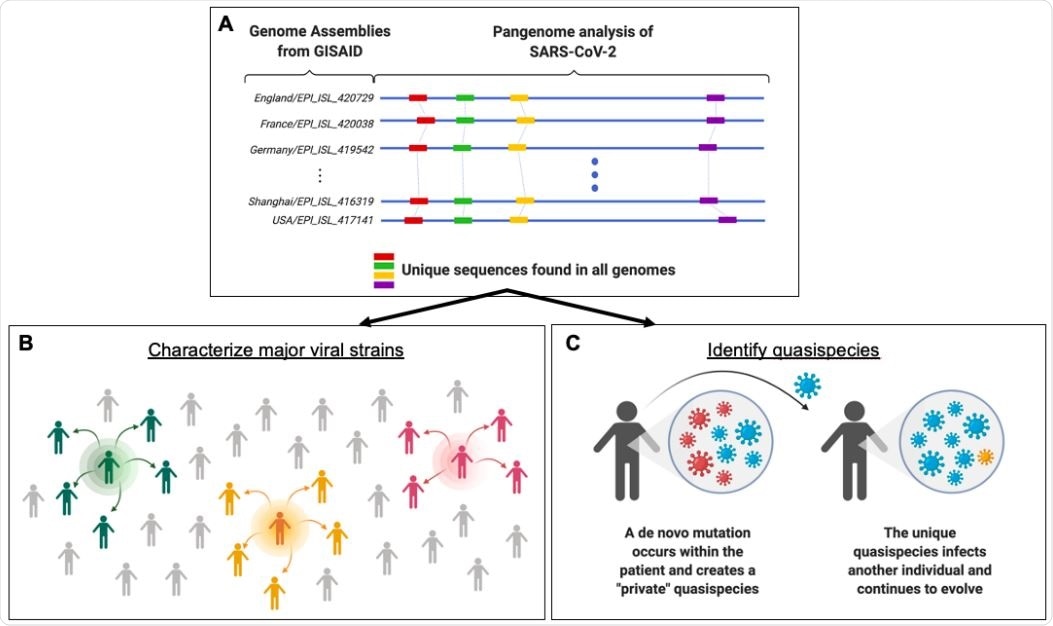
Study suggests SARS-CoV-2 mutation fingerprints could be used for molecular contact tracing
Ethical issues raised by new genomic technologies: the case study of newborn genome screening, Cambridge Prisms: Precision Medicine

Rami Mehio on LinkedIn: Video: Targeting the right drug, at the

Maria Martínez-Fresno Moreno on LinkedIn: National Rapid Genome Sequencing in Neonatal Intensive Care

Rosy Volpi on LinkedIn: Meet Illumina at ESHG 2023
Copy Number Variation (CNV) Analysis

Rosy Volpi on LinkedIn: Session 2: Expanding Frontiers of Genomic

James Han on LinkedIn: Technical spotlight: Detecting small- and
from
per adult (price varies by group size)






