Genotyping by low-coverage whole-genome sequencing in intercross
By A Mystery Man Writer
Description

PDF) Genotype imputation in F2 crosses of inbred lines

PDF) Low-coverage sequencing in a deep intercross of the Virginia body weight lines provides insight to the polygenic genetic architecture of growth: novel loci revealed by increased power and improved genome-coverage

Figure S2. Visualization of the number of crossovers in all F2

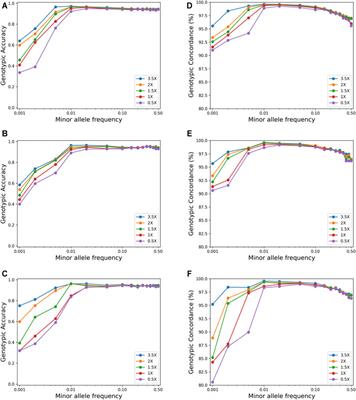
Assessment of the performance of different imputation methods for low-coverage sequencing in Holstein cattle - ScienceDirect
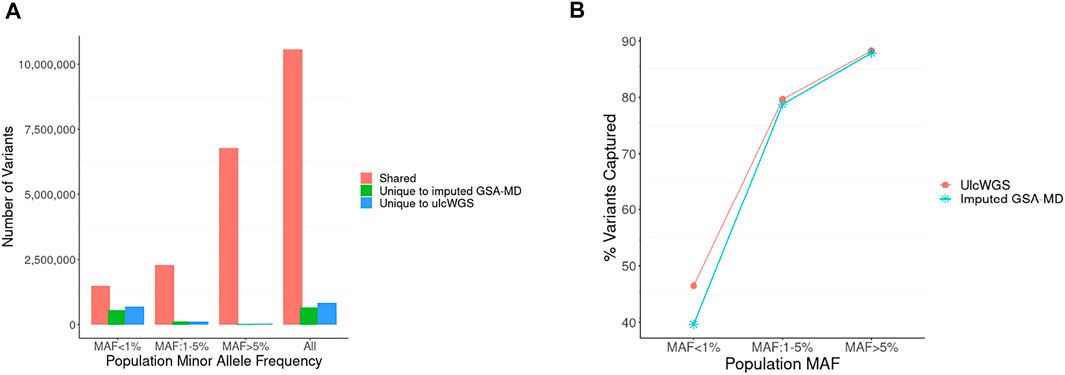
Frontiers Towards a Cost-Effective Implementation of Genomic Prediction Based on Low Coverage Whole Genome Sequencing in Dezhou Donkey

Assessment of the performance of different imputation methods for low-coverage sequencing in Holstein cattle - ScienceDirect

Whole-genome sequencing (WGS) and workflow of variant discovery. (A)
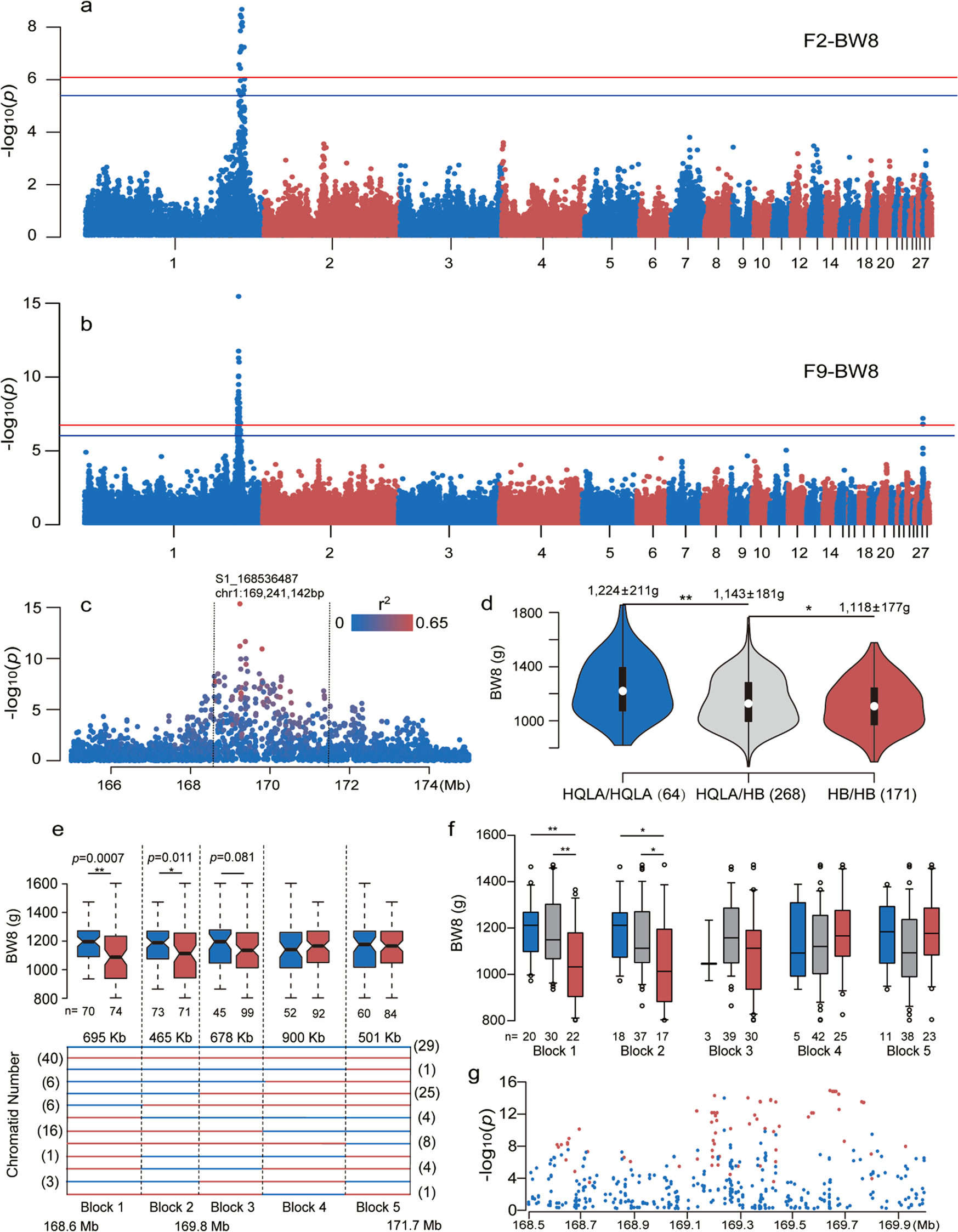
Multiple ancestral haplotypes harboring regulatory mutations cumulatively contribute to a QTL affecting chicken growth traits

Characterization of a haplotype-reference panel for genotyping by low-pass sequencing in Swiss Large White pigs, BMC Genomics

Genome-wide variant coverage and diversity. (a) Circos plot containing
from
per adult (price varies by group size)